How to Create AI Cell Signaling Pathway Diagrams
Three AI methods for publication-ready cell signaling pathway diagrams in minutes — text-to-figure, sketch-to-figure, and SVG vector export.
If you have ever spent an entire afternoon arranging protein nodes in Adobe Illustrator — nudging arrows by two pixels, hunting for a receptor tyrosine kinase clipart that doesn't look like it was drawn in 2003 — you already understand the problem. Cell signaling pathway diagrams are essential to virtually every molecular biology paper, yet producing one that meets journal standards can consume four to eight hours of skilled labor. BioRender offers a shortcut, but its subscription costs can easily exceed $1,000 per year for a single researcher, and the symbol library still forces you to work within rigid templates. There is a better way, and it does not require a design degree or an institutional budget.
The Old Way vs. The AI Way
| Step | Traditional | AI-Assisted |
|---|---|---|
| Initial draft | 2–3 hours | < 2 minutes |
| Revision cycle | 1–2 hours each | Seconds per iteration |
| Vector export | Manual cleanup | One-click SVG export |
| Skill required | Intermediate design | Plain-language prompting |
The gap is not incremental. It is the difference between a scientific figure being a bottleneck and a scientific figure being a routine deliverable.
Method 1 — Text-to-Figure (The Fastest Approach)
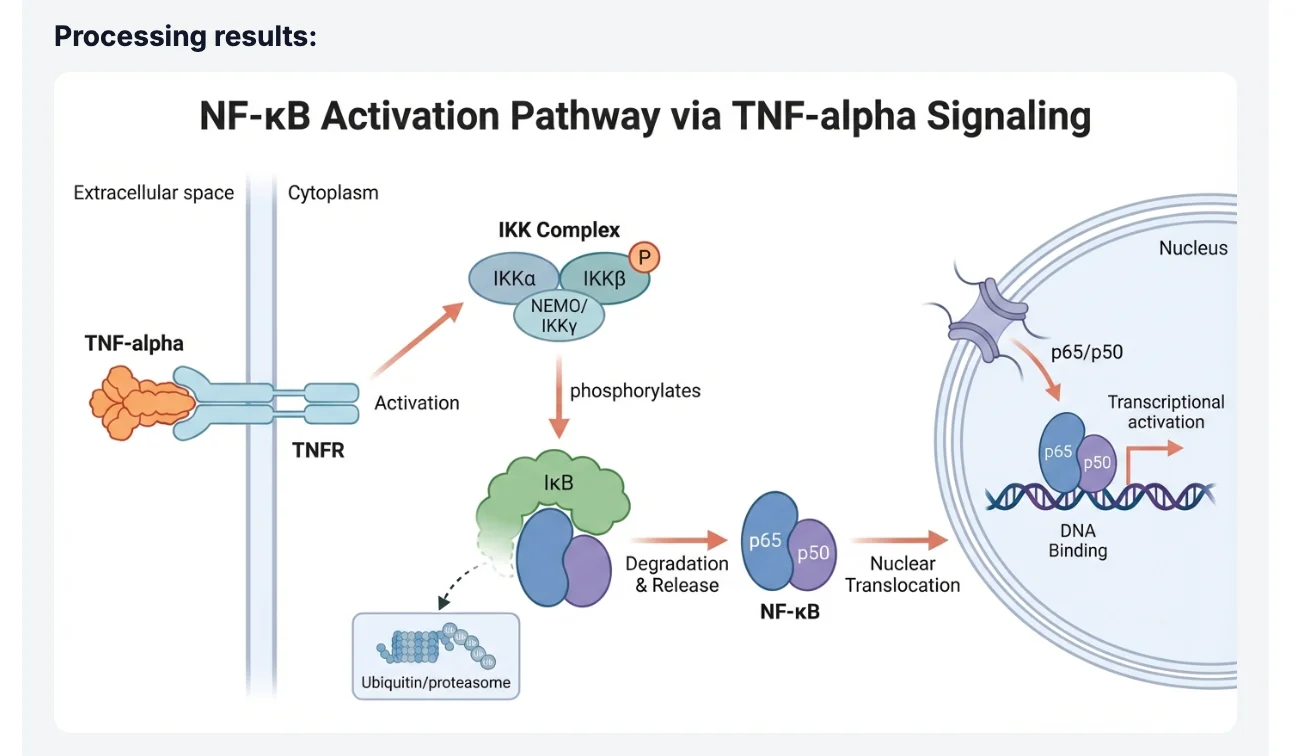
"Create a publication-ready cell signaling diagram of the canonical NF-κB pathway. Show TNF-α binding to TNFR1 at the plasma membrane, recruitment of TRADD and TRAF2, activation of the IKK complex (IKKα, IKKβ, IKKγ/NEMO), phosphorylation and proteasomal degradation of IκBα, and nuclear translocation of the p65/p50 heterodimer. Use a clean white background, labeled arrows indicating phosphorylation events, and a color scheme suitable for grayscale printing."
Notice what this prompt accomplishes: it names specific proteins rather than using generic terms, it specifies the subcellular compartments (plasma membrane, cytoplasm, nucleus), it requests labeled arrows for mechanistic clarity, and it anticipates a practical constraint (grayscale printing). Each of these details guides the model toward a scientifically accurate and journal-appropriate output.
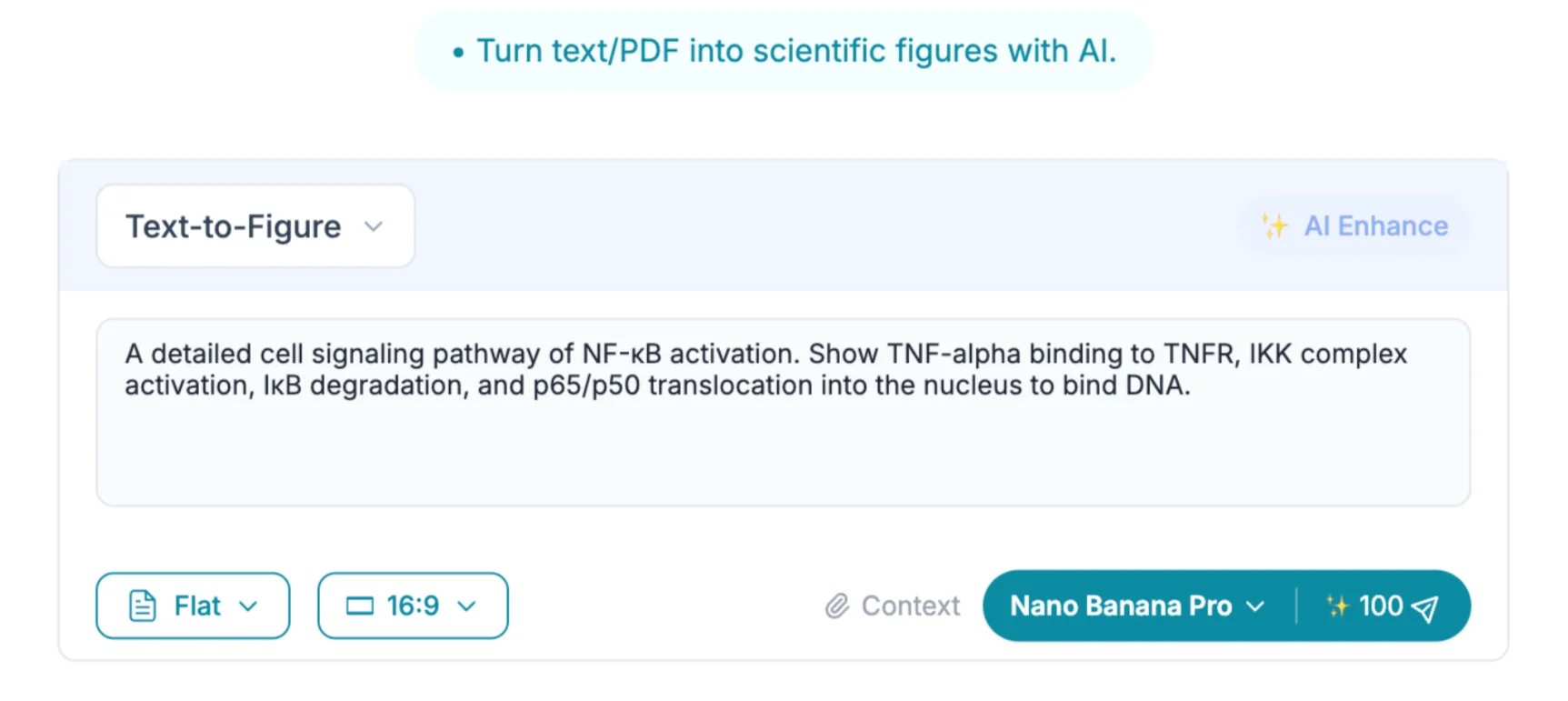
After submitting the prompt, the model generates a complete pathway diagram with consistent iconography, directional arrows, and protein labels. Most prompts produce a usable first draft; a single iteration — adding a detail like "include the p38 MAPK crosstalk pathway branching from TRAF2" — typically resolves any missing components.
Tip
See AI Scientific Figure Generation in Action
Watch how researchers create publication-ready scientific figures from text descriptions.
Explore the ToolMethod 2 — Image-to-Figure (From Sketch to Science)
The workflow is straightforward. Draw or photograph your sketch — it does not need to be clean; even pencil on a whiteboard qualifies — and upload it to the Image-to-Figure interface. Then add a short text prompt describing the style and any elements you want added or modified:
"Convert this hand-drawn MAPK cascade sketch into a publication-ready pathway diagram. Preserve the existing layout. Add labels for MEK1/2 and ERK1/2, use standard phosphorylation arrow notation, and apply a consistent blue-and-white color scheme."
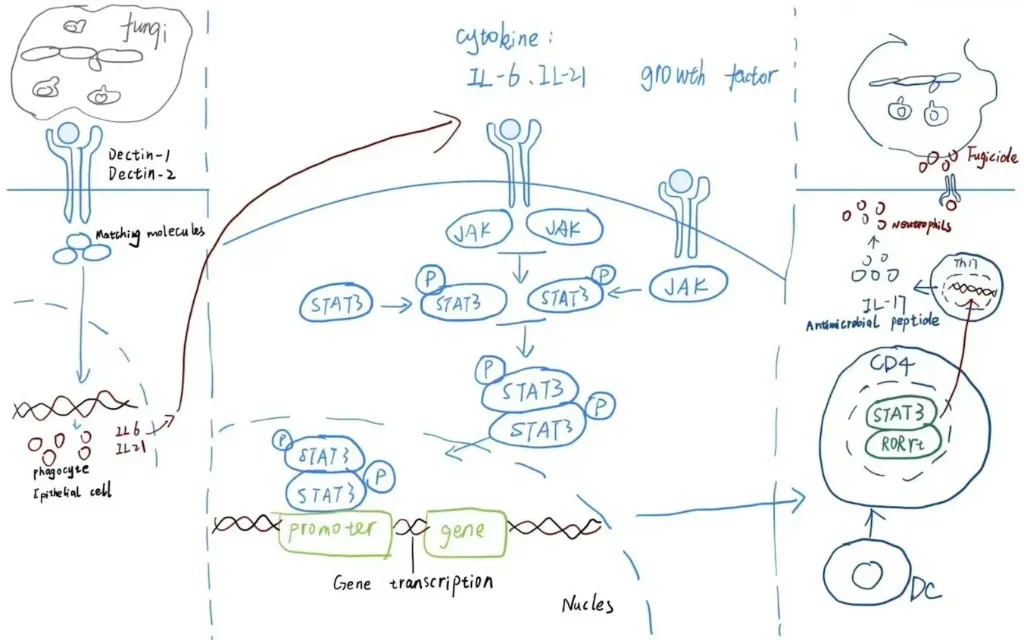
The model reads the spatial relationships in your sketch — which components are upstream, how branches connect, where the nucleus sits relative to the membrane — and renders a professional figure that honors your intended architecture while replacing rough linework with clean vector-style graphics.
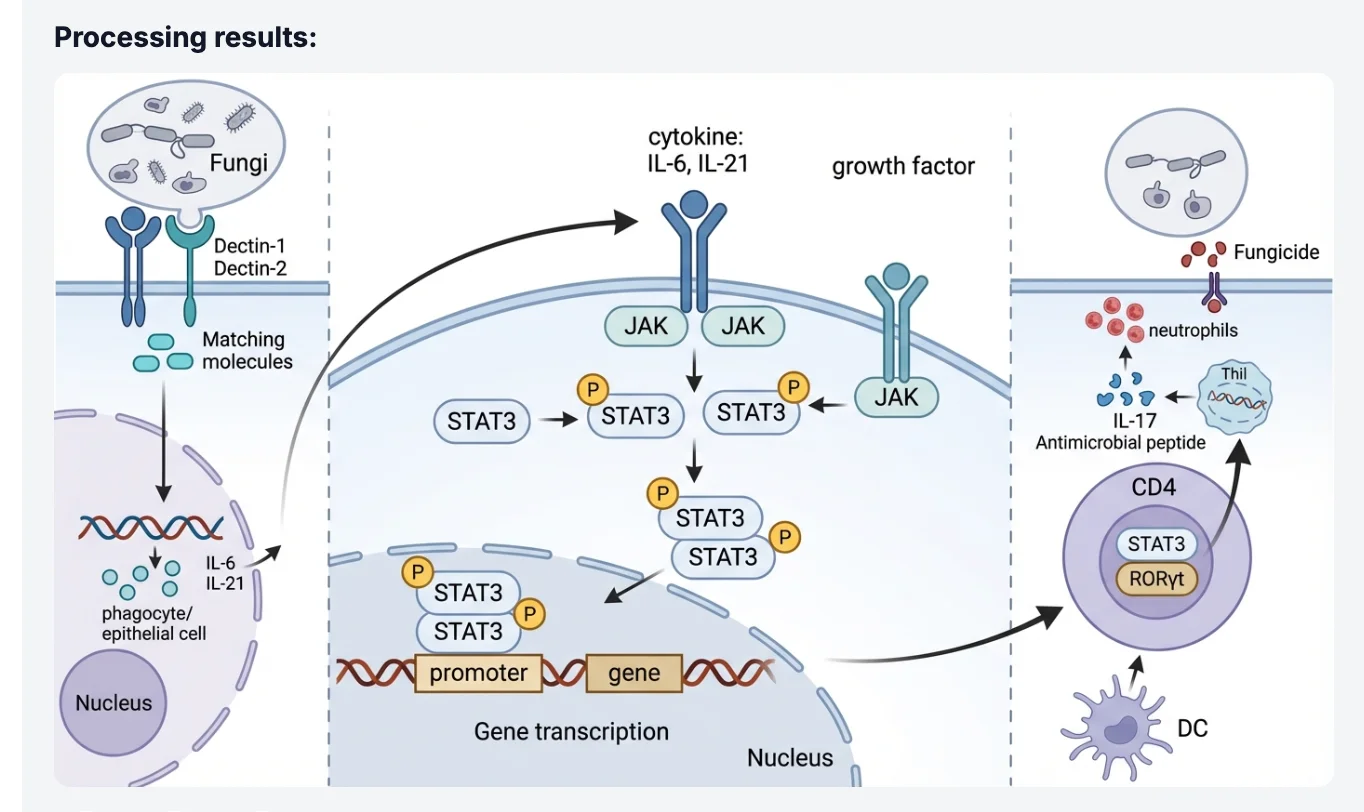
This approach is particularly valuable when you need to reproduce a pathway from a published paper in higher quality. Rather than redrawing from memory, you can photograph the original figure and instruct the AI to re-render it in your lab's style guide — saving time while maintaining scientific fidelity to the source.
The Secret Weapon — SVG Vectorizer
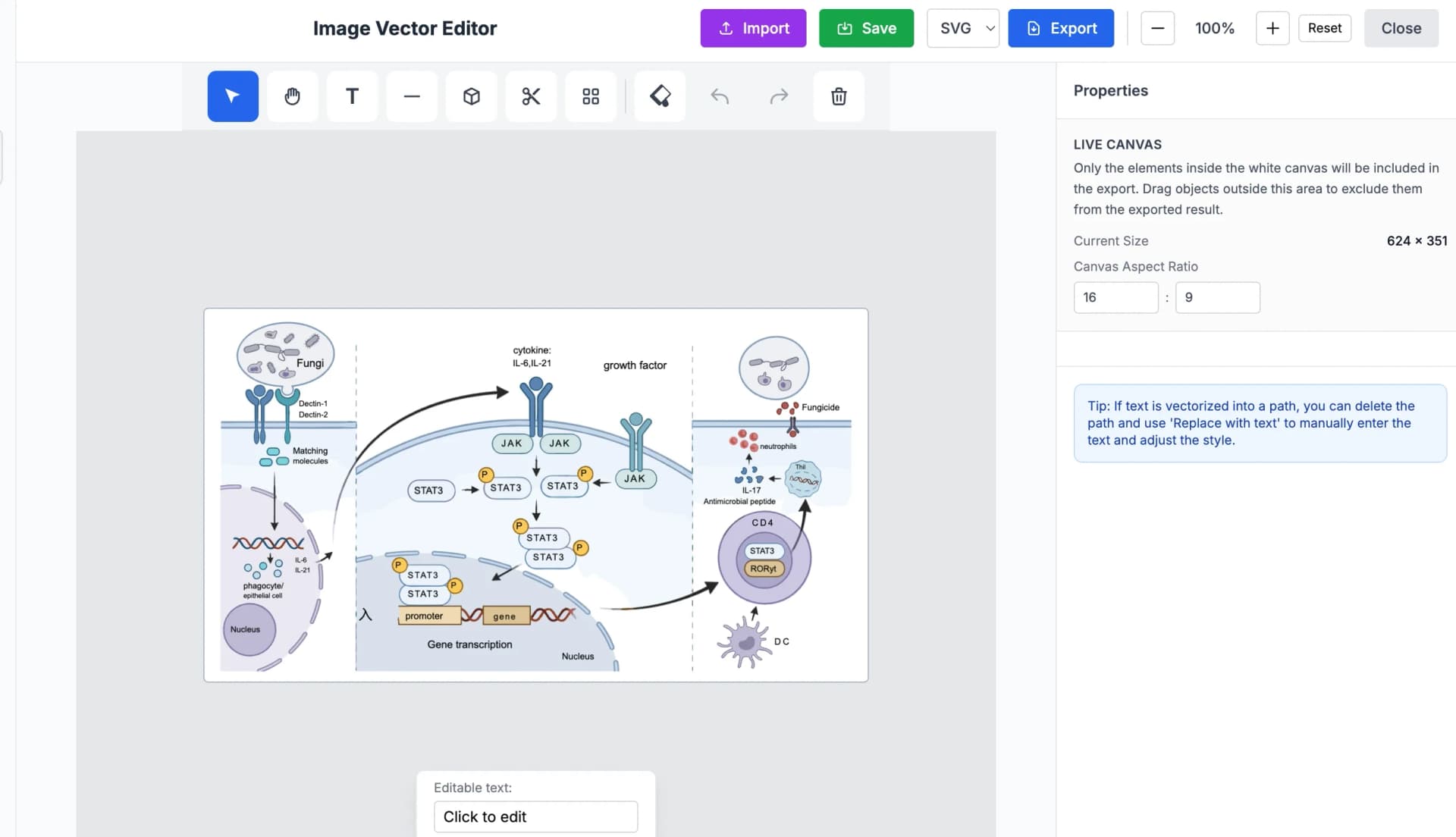
After generating a pathway diagram with either of the methods above, run it through the vectorization step. The tool traces every element — protein shapes, arrows, labels, background fills — and converts the raster output into a fully editable SVG file.
Once you have an SVG, you can open it in any vector editor (Inkscape, Adobe Illustrator, Affinity Designer) and manipulate individual components: change a protein label font, recolor a phosphorylation arrow, move a receptor without disturbing the rest of the scientific figure, or swap a single element for a revised version after peer review.
Create Scientific Figures Now
Describe your scientific figure in natural language — get publication-ready illustrations in minutes.
Try FreeTips for Better Pathway Diagrams
Generating a good pathway diagram is a skill that improves quickly with practice. Here are the most impactful adjustments researchers discover after their first few attempts: